Discussion
The PLA2 sequences found in C. s. tzabcan highly resembled the sequences of known crotoxin subunits. One of the sequences was 99% identical to crotoxin A from C. d. terrificus. The only difference between the two sequences was amino acid 93, where the sequence from C. s. tzabcan had glycine (G) and the PLA2 from C. d. terrificus had glutamic acid (E). However, sistruxin A from S. c. tergeminus also had glycine as its 93rd amino acid, suggesting that this difference does not change the function of the enzyme. The similarity of the sequences indicates that the protein from C. s. tzabcan would likely be structured in the same manner as crotoxin A, making it a PLA2. Due to the relationship between structure and function in PLA2s, this similarity also indicates that the PLA2 from C. s. tzabcan has an identical function to crotoxin A and other crotoxin A-like proteins, making it an amplifier of neurotoxicity. Another protein from C. s. tzabcan was found to have an amino acid sequence that was 100% identical to crotoxin B. This indicates that this sequence is likely a PLA2 with an identical structure and function to crotoxin B, making it neurotoxic. The presence of bands of protein from C. s. tzabcan’s venom in the PLA2 size range also indicates that there are PLA2s in C. s. tzabcan venom which would have the sequences that were found in this study. These findings also indicate that C. s. tzabcan has highly neurotoxic venom because sequences similar to both subunits of crotoxin were found.
One PLA2 sequence found in C. basiliscus was 100% identical to the sequence of crotoxin B found in C. d. terrificus. This indicates that this sequence is likely a PLA2 with the same structure and function as crotoxin B, making it neurotoxic. The C. basiliscus venom had a protein band in the PLA2 size range, suggesting that there are PLA2s in C. basiliscus venom which would have the sequence that was found. The presence of PLA2s with the identified sequence in C. basiliscus venom was also supported because the amino acid sequence found in this study was identical to the partial sequence that was found before. While a sequence resembling crotoxin B was found, no sequence was found from C. basiliscus which resembled crotoxin A. This could be because C. basiliscus venom is only slightly neurotoxic, or because not enough samples of C. basiliscus DNA were taken.
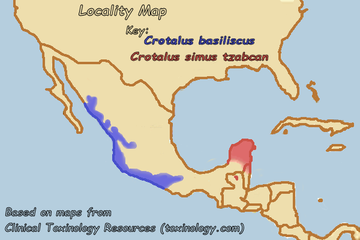
The crotoxin B-like sequences from C. s. tzabcan and C. basiliscus were also 100% identical. Since the ranges of C. s. tzabcan and C. basiliscus do not overlap (which is shown in the map below), it is unlikely that this similarity is due to a hybrid between the two species. This suggests that the two species have a common ancestor which also had this PLA2.
The similarity of the PLA2 sequences found in this study to the sequences of PLA2s with known functions supports the hypothesis that the function of PLA2s can be discovered by comparing their sequences to those of other PLA2s. In order to further test this hypothesis, it would be necessary to isolate the PLA2 proteins from the venoms of both snakes, sequence these proteins, and test their functions using assays.
Further research could also be done using the DNA sequences found in this study. These sequences could be used to manipulate the PLA2 genes, changing the amino acid sequences of the proteins. This could be used to determine which amino acids influence the function of PLA2 proteins and to learn more about the structure-function relationship in PLA2s. This would allow for a better understanding of the PLA2 protein, which could potentially lead to advances in many fields such as anti-venoms and drug delivery systems.
One PLA2 sequence found in C. basiliscus was 100% identical to the sequence of crotoxin B found in C. d. terrificus. This indicates that this sequence is likely a PLA2 with the same structure and function as crotoxin B, making it neurotoxic. The C. basiliscus venom had a protein band in the PLA2 size range, suggesting that there are PLA2s in C. basiliscus venom which would have the sequence that was found. The presence of PLA2s with the identified sequence in C. basiliscus venom was also supported because the amino acid sequence found in this study was identical to the partial sequence that was found before. While a sequence resembling crotoxin B was found, no sequence was found from C. basiliscus which resembled crotoxin A. This could be because C. basiliscus venom is only slightly neurotoxic, or because not enough samples of C. basiliscus DNA were taken.
The crotoxin B-like sequences from C. s. tzabcan and C. basiliscus were also 100% identical. Since the ranges of C. s. tzabcan and C. basiliscus do not overlap (which is shown in the map below), it is unlikely that this similarity is due to a hybrid between the two species. This suggests that the two species have a common ancestor which also had this PLA2.
The similarity of the PLA2 sequences found in this study to the sequences of PLA2s with known functions supports the hypothesis that the function of PLA2s can be discovered by comparing their sequences to those of other PLA2s. In order to further test this hypothesis, it would be necessary to isolate the PLA2 proteins from the venoms of both snakes, sequence these proteins, and test their functions using assays.
Further research could also be done using the DNA sequences found in this study. These sequences could be used to manipulate the PLA2 genes, changing the amino acid sequences of the proteins. This could be used to determine which amino acids influence the function of PLA2 proteins and to learn more about the structure-function relationship in PLA2s. This would allow for a better understanding of the PLA2 protein, which could potentially lead to advances in many fields such as anti-venoms and drug delivery systems.